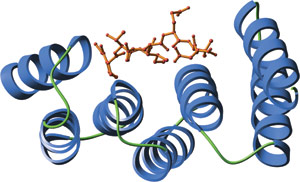
The amino-terminal TPR domain of human Hop bound to a carboxy-terminal peptide from Hsp70 (red).
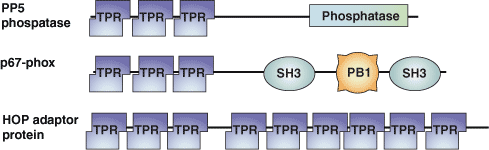
The tetratricopeptide repeat (TPR) motif was originally identified in yeast as a protein-protein interaction module in cell cycle proteins. It has since been found in organisms ranging from bacteria to humans. The TPR motif is a degenerate sequence of ~34 amino acids loosely based around the consensus residues -W-LG-Y-A-F-A-P-. The sequence occurs in tandem arrays and is present in over 800 different proteins. TPR motif-containing proteins act as scaffolds for the assembly of different multiprotein complexes including the anaphase promoting, the peroxisomal import receptor and the NADPH oxidase complexes.
TPR motifs exhibit a large degree of sequence diversity. However, structural comparison reveals a highly conserved three-dimensional structure. Individual TPR domains are composed of two anti-parallel α helices separated by a turn. Multiple TPR domains arrange at regular angles and form a right-handed super helix. This creates a groove with a large amount of surface area available for ligand binding.
| TPR Domain Proteins | Binding Partners | Peptide Ligands |
| PEX5 | PTS-1 target signal | S-K-L-COOH |
| Hop | Hsp70 - C-term heptapeptide Hsp90 - C-term pentapeptide | E-E-V-D-COOH E-E-V-D-COOH |
| p67phox | GTP-Rac | surface contacts |