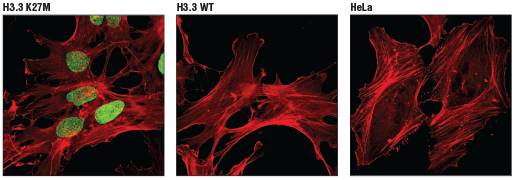
Oncohistones, mutated histones with oncogenic features, affect the global chromatin landscape and drive tumorigenesis due to altered gene expression. 78% of Diffuse Intrinsic Pontine Gliomas (DIPGs) contain a histone H3 oncohistone where a missense mutation substitutes Lys 27 with methionine, affecting its ability to be methylated and repress gene expression.
H3K27M serves as an oncohistone, where mutations contribute to tumor development because Ezh2 is no longer able to methylate the histone and gene expression is aberrantly upregulated.
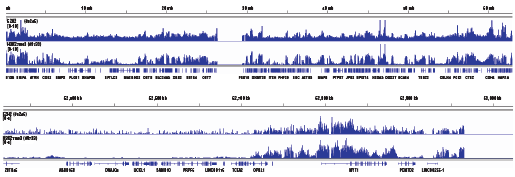
Changes in methylation of H3K27 are mostly due to mislocalization of Ezh2 on the genome and can be confirmed by ChIP-seq analysis.