| Cat. # | Size | Qty. | Price |
|---|---|---|---|
| 56593S | 100 µl |
|
| REACTIVITY | All |
| SENSITIVITY | Endogenous |
| MW (kDa) | |
| Source/Isotype | Rabbit IgG |
Product Information
This antibody has been shown by an independent laboratory to work in RNA-IP-seq. Please use at an assay-dependent dilution.
Note: This protocol is written for spotting either purified
total RNA or poly A-purified mRNA (titration of 2 μg, 1 μg, 500 ng,
250 ng, 125 ng, 62.5 ng, and 31.25 ng) onto a positively charged nylon
membrane using a 96-well dot blotting apparatus. Depending on the source of
the RNA, more or less RNA may be required for detection with the antibody.
Before Starting:
• RNA is sensitive to degradation by RNases, which can affect sample
integrity. It is recommended that all surfaces and equipment undergo RNase
decontamination.
• Purify total RNA and/or mRNA from cell pellet using an RNA isolation
kit. Assess total RNA quality by gel electrophoresis on a 1% agarose gel.
The 28S and 18S RNA should present as distinct bands. Smearing indicates RNA
degradation. See Figure 1.
• Cut a piece of nylon membrane to fit the size of the dot blot
manifold.
• Wet nylon membrane with 10X SSC Buffer.
• Dry membrane by placing it in a 96-well dot blot apparatus and
applying vacuum.
Optional: To normalize sample loading using methylene blue,
apply stain before Section C, Step 1 and capture an image. Rinse blots three
times for 5 min each with 15 mL dH2O. Stain does not affect
antibody binding or detection.
NOTE: Due to the kinetics of the detection reaction, signal is most intense immediately following incubation and declines over the following 2 hr.
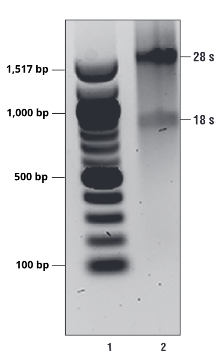
Figure 1. Representative image of isolated, intact total RNA. 28S and 18S RNA should migrate as distinct bands. If RNA presents as a smear, the sample may be degraded and unfit to use in downstream assays. Lane 1 is NEB 100 bp DNA ladder and lane 2 is total RNA isolated from 293T cells.
posted November 2018
Protocol Id: 1784
All Species Expected
Monoclonal antibody is produced by immunizing animals with N6-methyladenosine.
N6-methyladenosine (m6A) is a post-transcriptional modification found in various RNA subtypes. While the presence of m6A in RNA was described decades ago, the lack of tools has made interrogating the epitranscriptomic landscape challenging (1,2). With the emergence of new technologies such as miCLIP and NG-RNA-seq, researchers have been able to show that m6A is a biologically relevant mark in mRNA that is enriched in 3’ UTRs and stop codons (3,4). The m6A writer complex consists of a core heterodimer of methyltransferase-like protein 3 (METTL3) and methytransferase-like protein 14 (METTL14), and the additional regulatory proteins Virlizer/VIRMA and Wilms tumor 1-associated protein (WTAP) (5). METTL3 is the catalytic methyltransferase subunit and METTL14 is the target recognition subunit that binds to RNA (6). The Virilzer/VIRMA protein directs m6A methylation to the 3’ UTRs and stop codons, and WTAP targets the complex to nuclear speckles, which are sites of RNA processing (7). Less is known about readers and erasers of m6A, and while the fat mass and obesity-associated protein FTO was the first discovered m6A demethylase, subsequent studies demonstrated that this enzyme may prefer the closely related m6Am mark in vivo (8,9). ALKBH5 was later shown to be a bona fide m6A demethylase enzyme, contributing to the idea that the m6A modification is dynamically regulated (10). Readers of the m6A mark include the YTH protein family, which can bind to m6A and influence mRNA stability and translation efficiency (3,11-13). The m6A mark and machinery have been shown to regulate a variety of cellular functions, including RNA splicing, translational control, pluripotency and cell fate determination, neuronal function, and disease (1, 14-17). The m6A writer complex has been linked to various cancer types including AML and endometrial cancers (18,19). Additionally, m6A has been implicated in resistance to chemotherapy (20).
Except as otherwise expressly agreed in a writing signed by a legally authorized representative of CST, the following terms apply to Products provided by CST, its affiliates or its distributors. Any Customer's terms and conditions that are in addition to, or different from, those contained herein, unless separately accepted in writing by a legally authorized representative of CST, are rejected and are of no force or effect.
Products are labeled with For Research Use Only or a similar labeling statement and have not been approved, cleared, or licensed by the FDA or other regulatory foreign or domestic entity, for any purpose. Customer shall not use any Product for any diagnostic or therapeutic purpose, or otherwise in any manner that conflicts with its labeling statement. Products sold or licensed by CST are provided for Customer as the end-user and solely for research and development uses. Any use of Product for diagnostic, prophylactic or therapeutic purposes, or any purchase of Product for resale (alone or as a component) or other commercial purpose, requires a separate license from CST. Customer shall (a) not sell, license, loan, donate or otherwise transfer or make available any Product to any third party, whether alone or in combination with other materials, or use the Products to manufacture any commercial products, (b) not copy, modify, reverse engineer, decompile, disassemble or otherwise attempt to discover the underlying structure or technology of the Products, or use the Products for the purpose of developing any products or services that would compete with CST products or services, (c) not alter or remove from the Products any trademarks, trade names, logos, patent or copyright notices or markings, (d) use the Products solely in accordance with CST Product Terms of Sale and any applicable documentation, and (e) comply with any license, terms of service or similar agreement with respect to any third party products or services used by Customer in connection with the Products.