Scientists from the Site Discovery Group at Cell Signaling Technology ( CST ) recently published a paper in the journal Science Signaling ( Moritz, A. et al. 2010. Sci Signal. 3, ra64 ). The study aims to answer some of the most basic questions in cell biology—what drives a cancer cell to grow and survive? And, can we stop growth through the use of targeted inhibitors? Results from this study provide valuable information on the aberrant signaling mechanisms found in lung cancer, identifies potential therapeutic drug targets, and may help to predict therapies for patients with the disease.
Receptor tyrosine kinases ( RTKs ) are a class of signaling molecules that can drive the growth and survival of tumor cells in many forms of cancer, and are sometimes referred to as oncogenic drivers. Downstream of RTKs , there are several core signaling pathways commonly found to be responsible for cancer growth and survival; these include the PI3K/Akt, MAPK, and mTOR pathways. All of these pathways activate AGC kinase family members, including Akt, RSK, and p70 S6 kinases, whose protein substrates are phosphorylated at the RxRxxS/T motif. Previously, large-scale phosphoproteomic analysis of these serine/threonine kinases has not been performed due to technical challenges specific to arginine-rich motif sequences.
Listen to an interview with senior author, Dr. Michael Comb, as he talks about the results of this paper in a Science Signaling podcast.
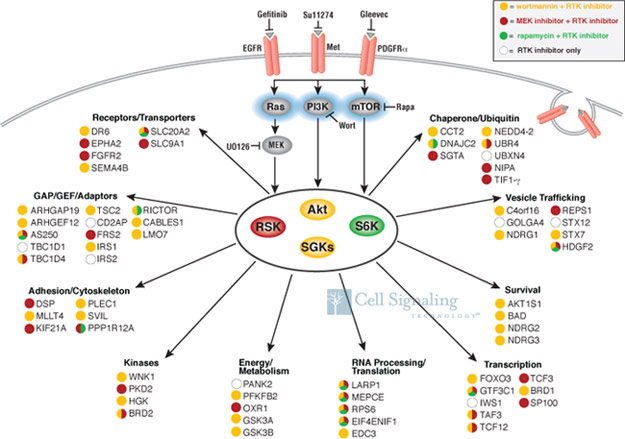
This study uses three previously characterized non-small cell lung cancer (NSCLC) cell lines in which cancer growth and survival is driven by one of three different RTKs : EGFR, c-Met, or PDGFR. The experimental approach involved the use of PhosphoScan®, CST ’s proprietary methodology for antibody-based peptide enrichment combined with tandem mass spectrometry for quantitative profiling of post-translational modifications. A key step was the development of a RxRxxS/T Motif Antibody, which was then used as an affinity reagent to selectively immunoprecipitate phosphorylated substrates of Akt, RSK, and p70 S6 kinases. Use of various RTK inhibitors in combination with inhibitors specific for PI3K, MEK, and mTOR allowed for mapping of the signaling network downstream of these RTKs . Over 300 known and novel substrates were identified, including proteins involved in scaffolding, protein stability, metabolism, trafficking, and motility (see figure below).
For more information, visit PhosphoSitePlus ® , CST ’s manually curated post-translational modification resource available at www.phosphosite.org , where all information on the observed substrate phosphorylation found in this study has been made available.