Immunofluorescence (IF) involves the labeling of cellular proteins with specific primary antibodies and fluorochrome-conjugated secondary antibodies (indirect method) or labeling with directly conjugated primary antibodies (direct method). Most fluorescence microscopes allow the examination of subcellular localization, relative expression level and/or activation-state (e.g. phosphorylation status) of two or three fluorescently-labeled proteins in a single sample. The ability to perform multiplexed analyses eliminates the need for consecutive sections when studying a subset of cells with multiple markers in a tissue, saving both time and reagents. Scientists at Cell Signaling Technology (CST) have validated over 800 activation-state specific (e.g., phosphorylation-specific) and total protein antibodies for IF applications, such as manual fluorescence microscopy or automated imaging and laser scanning high content platforms. All CST™ antibodies that are approved for use in immunofluorescent assays have undergone a rigorous validation process.
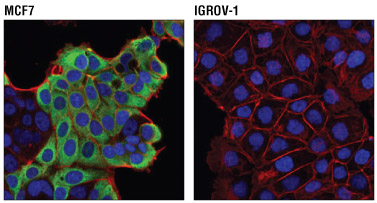
Confocal immunofluorescent analysis of MCF7 cells (positive, left) and IGROV-1 cells (negative, right) using AGR2 (D9V2F) XP® Rabbit mAb #13062 (green) and β-Actin (8H10D10) Mouse mAb #3700 (red). Blue pseudocolor= DRAQ5® #4084 (fluorescent DNA dye).
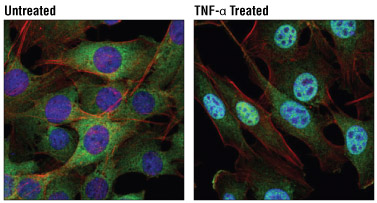
Confocal immunofluorescent analysis of C2C12 cells, untreated (left) or treated with Mouse Tumor Necrosis Factor-α (mTNF-α) #5178 (20 ng/ml, 30 min; right), using NF-κB1 p105/p50 (D4P4D) Rabbit mAb #13586 (green). Actin filaments were labeled with DyLight™ 554 Phalloidin #13054 (red). Blue pseudocolor= DRAQ5® #4084 (fluorescent DNA dye).
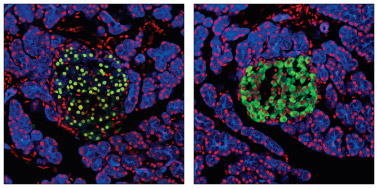
Pdx1 Antibody #2437: Confocal immunofluorescent analysis of normal rat pancreas using Pdx1 Antibody #2437 (green, left) or Insulin (C27C9) Rabbit mAb #3014 (green, right). Keratin filaments were labeled with Pan-Keratin (C11) Mouse mAb (Alexa Fluor® 647 Conjugate) #4528 (blue). Red = Propidium Iodide/RNase #4087 (fluorescent DNA dye).
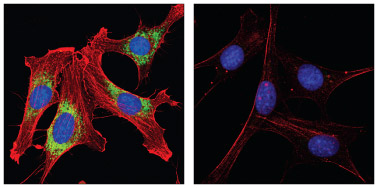
PDI Antibody #2446 and β-Actin (8H10D10) Mouse mAb #3700: Confocal IF analysis of NIH/3T3 cells, permeabilized with methanol (left) or 0.3% Triton X-100 (right), using #2446 (green) and #3700 (red). Blue pseudocolor = DRAQ5® #4084 (fluorescent DNA dye).