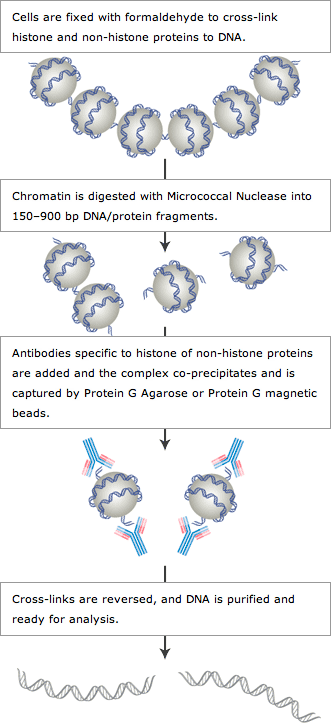
Overview of Chromatin IP Assay Methodology
ChIP Assay Overview
The chromatin immunoprecipitation (ChIP) assay is a powerful and versatile technique used for probing protein-DNA interactions within the natural chromatin context of the cell (1,2). This assay can be used to identify multiple proteins associated with a specific region of the genome, or the opposite, to identify the many regions of the genome associated with a particular protein (3–6). In addition, the ChIP assay can be used to define the spatial and temporal relationship of a particular protein-DNA interaction. For example, the ChIP assay can be used to determine the specific order of recruitment of various protein factors to a gene promoter or to “measure” the relative amount of a particular histone modification across an entire gene locus during gene activation (3,4). In addition to histone proteins, the ChIP assay can also be used to analyze binding of transcription factors, transcription co-factors, DNA replication factors and DNA repair proteins.
When performing the ChIP assay, cells are first fixed with formaldehyde, a reversible protein-DNA cross-linking agent that serves to fix or “preserve” the protein-DNA interactions occurring in the cell (see method overview) (1,2). Cells are then lysed and chromatin is harvested and fragmented using either sonication or enzymatic digestion. The chromatin is then subjected to immunoprecipitation using antibodies specific to a particular protein or histone modification. Any DNA sequences that are associated with the protein or histone modification of interest will co-precipitate as part of the cross-linked chromatin complex and the relative amount of that DNA sequence will be enriched by the immunoselection process. After immunoprecipitation, the protein-DNA cross-links are reversed and the DNA is purified. The enrichment of a particular DNA sequence or sequences can then be detected by a number of different methods.
Standard PCR methods are often employed to identify the DNA sequences or regions of the genome associated with a particular protein or histone modification (1,2). PCR is used to measure the relative abundance of a particular DNA sequence enriched by a protein-specific immunoprecipitation versus an immunoprecipitation with a non-specific antibody control. PCR products are run on an agarose or acrylamide gel to facilitate quantification, and the level of enrichment of the DNA sequence is determined relative to the total amount of input DNA (percent of input). The level of enrichment can also be expressed as fold enrichment above background (enrichment relative to that of the non-specific antibody control). Real-Time PCR provides a more accurate, gel-free system for the quantification of DNA enrichment. Alternatively, the ChIP assay can be combined with genomic tiling micro-array (ChIP on chip) techniques, sequencing, or cloning strategies, which allow for genome-wide analysis of protein-DNA interactions and histone modifications (5–8).
Our enzymatic ChIP Kit for cells and tissues contains buffers and reagents needed to perform the ChIP assay with mammalian cells and works for both histone modifications and non-histone DNA-binding proteins. After cell lysis, the chromatin is fragmented by partial digestion with Micrococcal Nuclease to obtain chromatin fragments of 1 to 5 nucleosomes in size. Enzymatic digestion of chromatin is much milder than sonication and eliminates problems due to variability in sonication power and emulsification of chromatin during sonication, which can result in incomplete fragmentation of chromatin or loss of antibody epitopes due to protein denaturation and degradation. The chromatin immunoprecipitations are performed using antibodies and either ChIP Grade Protein G Agarose or ChIP Grade Protein G Magnetic Beads. After reversal of protein-DNA cross-links, the DNA is purified using DNA purification spin columns provided in the kit. The DNA purification spin columns combine the convenience of spin-column technology with the selective binding properties of a uniquely designed silica membrane that allows for efficient recovery of DNA and removal of protein contaminants without the need for phenol/chloroform extractions and ethanol precipitations. After DNA purification, the enrichment of particular DNA sequences can be analyzed by a variety of methods, including standard PCR, quantitative real-time PCR, amplification for ChIP on chip, sequencing or cloning techniques.
Method Overview
In addition to providing buffers and reagents required to perform the ChIP assay, the SimpleChIP® Kit provides important controls that allow for user determination of a successful ChIP experiment. The kit contains a positive control Histone H3 Antibody, a negative control Normal Rabbit IgG Antibody and primer sets for PCR detection of the ribosomal protein L30 (RPL30) gene locus (human and mouse primer sets included). Histone H3 is a core component of chromatin in the cell and is bound to most DNA sequences throughout the genome, including the RPL30 locus. Thus, immunoprecipitation of chromatin with the Histone H3 antibody will enrich for the RPL30 gene, while immunoprecipitation with the Normal Rabbit IgG will not result in RPL30 gene enrichment. This enrichment can be quantified using either standard PCR or quantitative real-time PCR methods and the RPL30 primer sets provided in the kit. Importantly, since histone H3 is bound to most DNA sequences throughout the genome, the Histone H3 Antibody serves as a positive control IP for almost any locus studied, giving the user even more confidence that their ChIP experiment was performed successfully.
The SimpleChIP® Kit provides enough reagents to perform up to 6 chromatin preparations (or optimizations) and 30 immunoprecipitations and is optimized for 4 X 107 cells per experiment. A ChIP assay can be performed in as little as two days and can easily be scaled up or down for use with more or fewer cells.
Background References
- Orlando V (2000) Mapping chromosomal proteins in vivo by formaldehyde-crosslinked-chromatin immunoprecipitation. Trends Biochem. Sci. 25(3), 99–104.
- Kuo MH, Allis CD (1999) In vivo cross-linking and immunoprecipitation for studying dynamic Protein:DNA associations in a chromatin environment. Methods 19(3), 425–33.
- Agalioti T, Lomvardas S, Parekh B, Yie J, Maniatis T, Thanos D (2000) Ordered recruitment of chromatin modifying and general transcription factors to the IFN-beta promoter. Cell 103(4), 667–78.
- Soutoglou E, Talianidis I (2002) Coordination of PIC assembly and chromatin remodeling during differentiation-induced gene activation. Science 295(5561), 1901–4.
- Mikkelsen TS, Ku M, Jaffe DB, Issac B, Lieberman E, Giannoukos G, Alvarez P, Brockman W, Kim TK, Koche RP, Lee W, Mendenhall E, O'Donovan A, Presser A, Russ C, Xie X, Meissner A, Wernig M, Jaenisch R, Nusbaum C, Lander ES, Bernstein BE (2007) Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448(7153), 553–60.
- Lee TI, Jenner RG, Boyer LA, Guenther MG, Levine SS, Kumar RM, Chevalier B, Johnstone SE, Cole MF, Isono K, Koseki H, Fuchikami T, Abe K, Murray HL, Zucker JP, Yuan B, Bell GW, Herbolsheimer E, Hannett NM, Sun K, Odom DT, Otte AP, Volkert TL, Bartel DP, Melton DA, Gifford DK, Jaenisch R, Young RA (2006) Control of developmental regulators by Polycomb in human embryonic stem cells. Cell 125(2), 301–13.
- Weinmann AS, Farnham PJ (2002) Identification of unknown target genes of human transcription factors using chromatin immunoprecipitation. Methods 26(1), 37–47.
- Wells J, Farnham PJ (2002) Characterizing transcription factor binding sites using formaldehyde crosslinking and immunoprecipitation. Methods 26(1), 48–56.