Benefits of Enzyme-based Chromatin Digestion
New SimpleChIP® Enzymatic Chromatin IP Kits from Cell Signaling Technology (CST) are co-developed by CST and NEB scientists and therefore contain the highest quality research reagents. These kits are available with either Protein G agarose or Protein G magnetic beads and contain all buffers and reagents needed to perform up to 30 ChIP assays. Reagents supplied in the kits are the product of in-house ChIP validation at CST, simplifying your optimization. The kits can be utilized with any ChIP-validated antibody to detect endogenous levels of protein-DNA interactions and histone modifications in mammalian cells. Protein G agarose and Protein G magnetic beads are also available as separate products.
Major Advantages of MNase Enzymatic Digestion Compared to Sonication:
- Enzymatic digestion is milder than sonication and better preserves the integrity of the chromatin and antibody epitopes, which means increased IP efficiency.
- Increased IP efficiency of target proteins means enhanced detection of protein-bound DNA loci.
- Enhanced detection of target DNA loci means less input chromatin required, saving valuable time and reagents.
- Enzymatic digestion provides greater sensitivity with lower background, which is essential when detecting DNA bound transcription factors and cofactors.
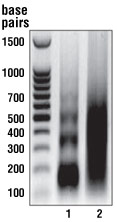
Enzyme-based and sonication-based ChIP kits produce chromatin fragments of a similar size. Chromatin was prepared from 4 x 107 HCT116 human colorectal carcinoma cells according to the protocols included with the SimpleChIP® Enzymatic Chromatin IP Kit (Magnetic Beads) #9003 or a competitor’s sonication-based ChIP kit. DNA was purified from each chromatin sample and DNA fragment size was determined by electrophoresis on a 1% agarose gel. Both enzymatic digestion with the SimpleChIP® Kit (lane 1) and sonication with the competitor’s kit (lane 2) produced chromatin fragments ranging from 150 to 700 bp, corresponding to one to five nucleosomes in length.
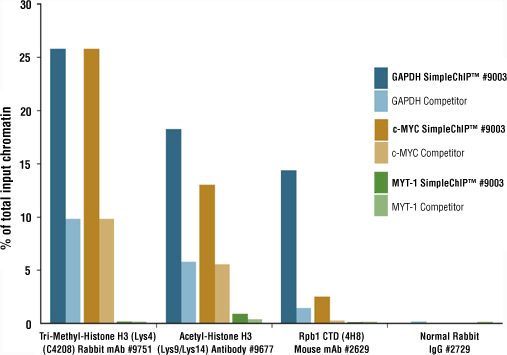
Example A: PCR primers are specific for the transcriptionally active GAPDH and c-MYC genes and the inactive MYT-1 gene.
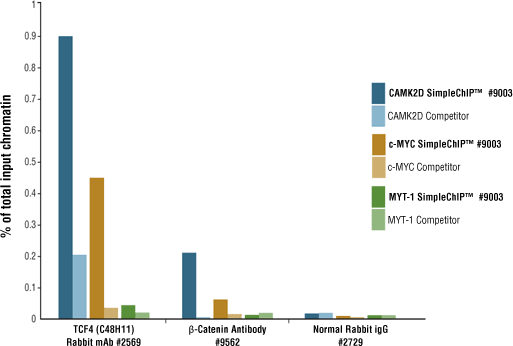
Example B: PCR primers are specific for known TCF4 binding sites in the CAMK2D and c-MYC genes, and a region of the MYT-1 gene that does not contain a TCF4 binding site.
SimpleChIP® digested chromatin is more conducive to immunoprecipitation than sonicated chromatin. Chromatin immunoprecipitations were performed with 10 μg of cross-linked HCT116 chromatin and the indicated antibodies, using SimpleChIP® Enzymatic Chromatin IP Kit (Magnetic Beads) #9003 or a competitor’s sonication-based ChIP Kit. The enriched DNA was quantified by qPCR. The amount of immunoprecipitated DNA in each sample is presented as a percent of the total input chromatin. For every target tested, enzyme-digested chromatin showed better enrichment of target DNA loci than did sonicated chromatin.